TADKit - Interactive Anomaly Detection demonstrator
import os
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from ouagen import ornstein_uhlenbeck_anomaly
t = np.arange(1000)
t = t / t.shape[0]
pre_X, pre_y = ornstein_uhlenbeck_anomaly(
t,
size=20,
noise_scale=1,
mean_reverting=3,
anomaly_freq=.1,
anomaly_duration=0.005,
anomaly_scale=40,
)
t = pd.to_datetime('2021-01-01') + pd.Timedelta(1, 'h') * np.arange(t.shape[0])
X = pd.DataFrame(pre_X, index=t, columns=[f'X{i}' for i in range(pre_X.shape[1])])
y = pd.DataFrame(pre_y[:, None], index=t, columns=['y'])
async_X = X.melt(value_vars=X.columns.tolist(), ignore_index=False).rename(
columns={"variable": "sensor", "value": "data"}
)
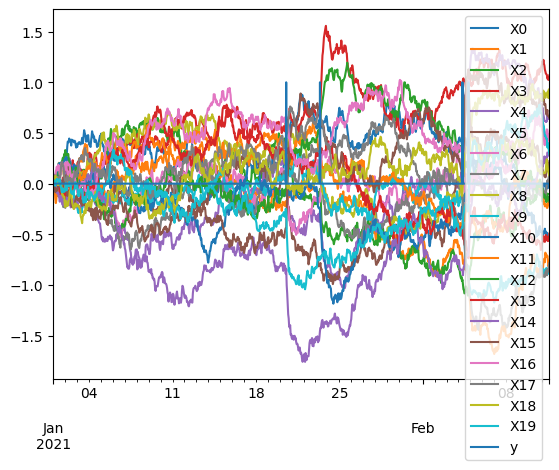
oua = pd.concat([X, y], axis=1)
oua.plot()
# oua.to_csv("oua.csv")
import ipywidgets as widgets
# Create a dropdown selection widget
select = widgets.Dropdown(
options=["synchronous", "asynchronous"],
value="synchronous",
description='Select:'
)
display(select)
from tadkit.catalog.rawtowideformatter import RawToWideFormatter
target_X = X if select.value == "synchronous" else async_X
formatter = RawToWideFormatter(
data=target_X,
backend="pandas"
)
print(f"Formatter has {formatter.available_properties=}")
Formatter has formatter.available_properties=['synchronous', 'pandas_backend', 'fixed_time_step']
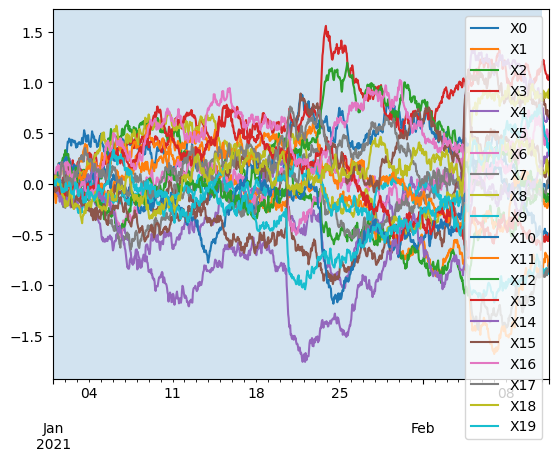
This allows to select a training interval, a resampling resolution and the set of target sensors for learning.
from query_selection import query_widget_selection
query_dict = query_widget_selection(formatter.query_description)
query_translated = {key: widget.value for key, widget in query_dict.items()}
X_train = formatter.format(**query_translated)
query_translated.pop("target_period")
X_test = formatter.format(**query_translated)
ax = X_test.plot()
ax.axvspan(X_train.index[0], X_train.index[-1], alpha=0.2)
plt.show()
from tadkit.base.registry import registry
import tadkit.catalog.registry_init # ensure registrations happen
registry.print_catalog_classes()
registry.list_learners()
registry.match_learners(formatter)
[registry] Skipping optional learner 'DataReconstructionAD': No module named 'cnndrad'
[registry] Skipping optional learner 'DiLAnoDetectm': No module named 'sbad_fnn'
[TADkit registered Catalog]
Class learner_name='IsolationForest' is registered in TADKit.
IsolationForest is operational in this environment.
IsolationForest is implicit child of TADLearner.
Class learner_name='KDEOutlierDetector' is registered in TADKit.
KDEOutlierDetector is operational in this environment.
KDEOutlierDetector is implicit child of TADLearner.
Class learner_name='GMMOutlierDetector' is registered in TADKit.
GMMOutlierDetector is operational in this environment.
GMMOutlierDetector is implicit child of TADLearner.
Class learner_name='TopologicalAnomalyDetector' is registered in TADKit.
TopologicalAnomalyDetector is operational in this environment.
TopologicalAnomalyDetector is implicit child of TADLearner.
Class learner_name='KcpLearner' is registered in TADKit.
KcpLearner is operational in this environment.
KcpLearner is implicit child of TADLearner.
[sklearn.ensemble._iforest.IsolationForest,
tadkit.catalog.sklearners.KDEOutlierDetector,
tadkit.catalog.sklearners.GMMOutlierDetector,
tdaad.anomaly_detectors.TopologicalAnomalyDetector,
kcpdi.kcp_ss_learner.KcpLearner]
for learner_cls in registry.match_learners(formatter):
learner = learner_cls() # instantiate directly
print(f"Instantiated {learner_cls.__name__}")
Instantiated IsolationForest
Instantiated KDEOutlierDetector
Instantiated GMMOutlierDetector
Instantiated TopologicalAnomalyDetector
Instantiated KcpLearner
from tadkit.utils.param_spec import params_from_class
from tadkit.utils.render_widgets_from_params import render_widgets_from_params
learner_params = {}
for learner_class in registry.match_learners(formatter):
learner_params.setdefault(learner_class.__name__, {})
print(learner_class.__name__)
params_specs = params_from_class(learner_class)
widgets, learner_params[learner_class.__name__] = render_widgets_from_params(params_specs)
IsolationForest
KDEOutlierDetector
GMMOutlierDetector
TopologicalAnomalyDetector
KcpLearner
learner_params["TopologicalAnomalyDetector"]()
{'step': 5,
'tda_max_dim': 1,
'contamination': 0.1,
'n_centers_by_dim': 5,
'random_state': 42,
'window_size': 20,
'support_fraction': None}
learners = {
learner_class.__name__:
learner_class(**learner_params[learner_class.__name__]())
for learner_class in registry.match_learners(formatter)
}
learners
{'IsolationForest': IsolationForest(),
'KDEOutlierDetector': KDEOutlierDetector(),
'GMMOutlierDetector': GMMOutlierDetector(),
'TopologicalAnomalyDetector': TopologicalAnomalyDetector(window_size=20),
'KcpLearner': KcpLearner()}
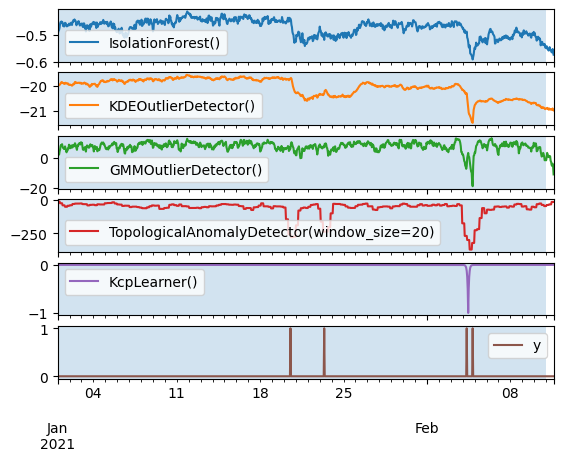
Fit on train query, score on the entire series.
# Fit loop:
anomalies = pd.DataFrame(index=X_test.index)
predictions = pd.DataFrame(index=X_test.index)
for learner_name, learner_object in learners.items():
print(f"Fitting {learner_name}")
learner_object.fit(X_train)
# Score loop:
for learner_name, fitted_learner_object in learners.items():
print(f"Scoring with {learner_name}")
anom_score = fitted_learner_object.score_samples(X_test)
anomalies[str(fitted_learner_object)] = anom_score
predictions[str(fitted_learner_object)] = fitted_learner_object.predict(X_test)
Fitting IsolationForest
Fitting KDEOutlierDetector
Fitting GMMOutlierDetector
Fitting TopologicalAnomalyDetector
Fitting KcpLearner
Scoring with IsolationForest
Scoring with KDEOutlierDetector
Scoring with GMMOutlierDetector
Scoring with TopologicalAnomalyDetector
Scoring with KcpLearner
axes = pd.concat([anomalies, y], axis=1).plot(subplots=True)
[ax.axvspan(X_train.index[0], X_train.index[-1], alpha=0.2) for ax in axes]
plt.show()
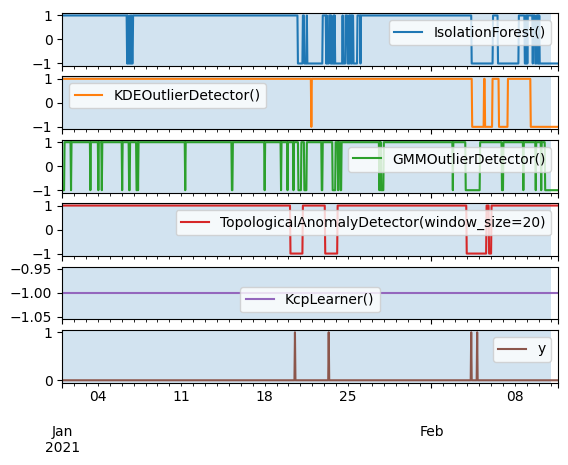
axes = pd.concat([predictions, y], axis=1).plot(subplots=True)
[ax.axvspan(X_train.index[0], X_train.index[-1], alpha=0.2) for ax in axes]
plt.show()
from plotly.offline import init_notebook_mode
from plotly.graph_objs import *
init_notebook_mode(connected=True) # initiate notebook for offline plot
pd.options.plotting.backend = "plotly"
df = pd.concat([anomalies.apply(lambda x: (x - x.min()) / (x.max() - x.min())), y], axis=1)
import plotly.express as px
fig = df.plot(color_discrete_sequence=px.colors.qualitative.Set2)
fig.update_layout(
legend=dict(
orientation="v",
entrywidth=100,
yanchor="bottom",
y=1.02,
xanchor="right",
x=1
),
width=1000,
height=600,
)
fig.show()
pd.options.plotting.backend = "matplotlib"